
Md. Helal Uddin Chowdhury* , Tuhin Das2 and Suranjana Sikdar2
Received: 2023-02-05 | Revised:2023-02-22 | Accepted: 2023-02-26 | Published: 2023-02-26
Abstract
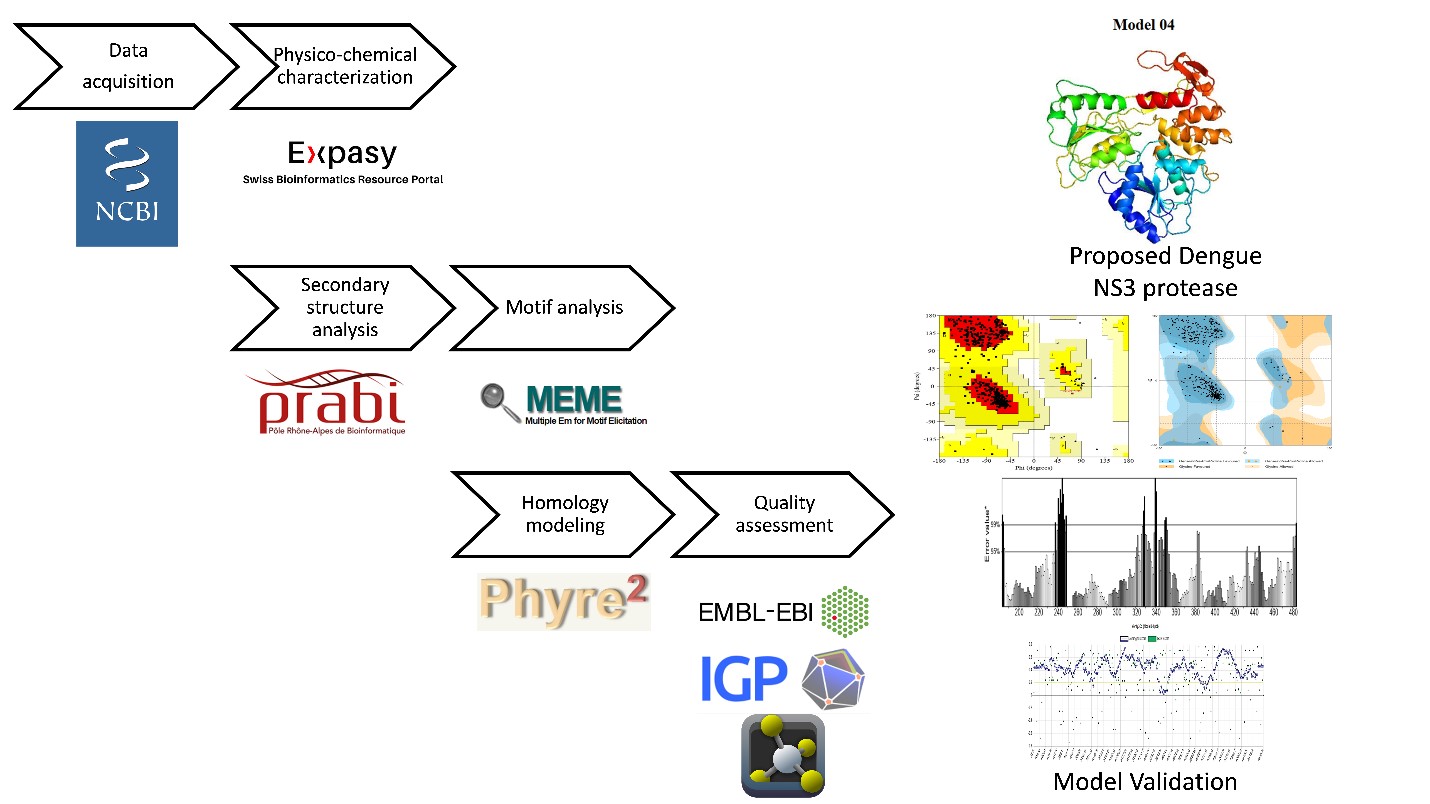
Dengue is causing significant morbidity and mortality worldwide. In poor and underdeveloped countries, the disease is spreading at an alarming rate due to a rise in population density and a decline in environmental cleanliness. Due to the mutation and variety of distinct dengue virus species, the disease is difficult to cure with standard techniques. In addition, there is still a need for effective vaccination against this fatal virus. Designing a vaccine needs a full explanation of the structural characteristics of the NS3 protease, the primary antigenic component of the virus. Several bioinformatics methods were utilized in this study to characterize the NS3 protease of the dengue virus utilizing data from various public databases. Different physio-chemical properties were determined using the ProtParam tool. Secondary structure and motifs were predicted using the SOPMA server and MEME suit. Finally, homology modeling of the selected protein was conducted using the PHYRE2 server. Quality assessment of the predicted structures was performed by employing Ramachandran plot, ERRAT, RAMPAGE, verify 3D, and RMSD scores to establish and suggest one best model for further experimentation. A satisfactory validation score in all those quality assessments implies the proposed model to be a good fit for the future experiment on this protein. Such homology modeling of the viral protein paves the way to a successful protein model and consequently leads to efficient vaccine design.
Keywords
Dengue 4 NS3 protease, Motif analysis, homology modeling
References
1. Samal, R.R.; Gupta, S.; Kumar, S. An overview of factors affecting dengue transmission in asian region and its predictive models. J. Appl. Nat. Sci. 2020, 12, 460470, doi:10.31018/jans.v12i3.2360.
2. Ramos-Castañeda, J.; Barreto dos Santos, F.; Martínez-Vega, R.; Galvão de Araujo, J.M.; Joint, G.; Sarti, E. Dengue in Latin America: Systematic Review of Molecular Epidemiological Trends. PLoS Negl. Trop. Dis. 2017, 11, e0005224, doi:10.1371/journal.pntd.0005224.
3. Ferreira-de-Lima, V.H.; Lima-Camara, T.N. Natural vertical transmission of dengue virus in Aedes aegypti and Aedes albopictus: a systematic review. Parasit. Vectors. 2018, 11, 77, doi:10.1186/s13071-018-2643-9.
4. Fink, A.L.; Williams, K.L.; Harris, E.; Alvine, T.D.; Henderson, T.; Schiltz, J.; Nilles, M.L.; Bradley, D.S. Dengue virus specific IgY provides protection following lethal dengue virus challenge and is neutralizing in the absence of inducing antibody dependent enhancement. PLoS Negl. Trop. Dis. 2017, 11, e0005721, doi:10.1371/journal.pntd.0005721.
5. Silva, N.M.; Santos, N.C.; Martins, I.C. Dengue and Zika Viruses: Epidemiological History, Potential Therapies, and Promising Vaccines. Trop. Med. Infect. Dis. 2020, 5, 150, doi:10.3390/tropicalmed5040150.
6. Uno, N.; Ross, T.M. Dengue virus and the host innate immune response. Emerg. Microbes Infect. 2018, 7, 1–11, doi:10.1038/s41426-018-0168-0.
7. Nasar, S.; Rashid, N.; Iftikhar, S. Dengue proteins with their role in pathogenesis, and strategies for developing an effective anti‐dengue treatment: A review. J. Med. Virol. 2020, 92, 941–955, doi:10.1002/jmv.25646.
8. Xu, T.; Sampath, A.; Chao, A.; Wen, D.; Nanao, M.; Chene, P.; Vasudevan, S.G.; Lescar, J. Structure of the Dengue virus helicase/nucleoside triphosphatase catalytic domain at a resolution of 2.4 A. J. Virol. 2005, 79, 10278–10288.
9. Norazharuddin, H.; Lai, N.S. Roles and Prospects of Dengue Virus Nonstructural Proteins as Antiviral Targets: An Easy Digest. Malays. J. Med. Sci. 2018, 25, 6–15, doi:10.21315/mjms2018.25.5.2.
10. Luo, D.; Vasudevan, S.G.; Lescar, J. The flavivirus NS2B–NS3 protease–helicase as a target for antiviral drug development. Antivir. Res. 2015, 118, 148–158, doi:10.1016/j.antiviral.2015.03.014.
11. Chowdhury, H.U.; Adnan, M.; Oh, K.K.; Cho, D.H. Decrypting molecular mechanism insight of Phyllanthus emblica L. fruit in the treatment of type 2 diabetes mellitus by network pharmacology. Phytomedicine Plus 2021, 100144, doi:10.1016/J.PHYPLU.2021.100144.
12. Adnan, M.; Jeon, B.B.; Chowdhury, M.H.U.; Oh, K.K.; Das, T.; Chy, M.N.U.; Cho, D.H. Network Pharmacology Study to Reveal the Potentiality of a Methanol Extract of Caesalpinia sappan L. Wood against Type-2 Diabetes Mellitus. Life. 2022, 12, 277, doi:10.3390/LIFE12020277.
13. Shovo, M.A.R.B.; Tona, M.R.; Mouah, J.; Islam, F.; Chowdhury, M.H.U.; Das, T.; Paul, A.; Ağagündüz, D.; Rahman, M.M.; Emran, T. Bin; et al. Computational and Pharmacological Studies on the Antioxidant, Thrombolytic, Anti-Inflammatory, and Analgesic Activity of Molineria capitulata. Curr. Issues Mol. Biol. 2021, 43, 434–456, doi:10.3390/CIMB43020035.
14. Enam, T.; Agarwala, R.; Chowdhury, M.H.U.; Chy, M.N.U.; Adnan, M. An in silico ADME/T and molecular docking studies of phytochemicals derived from Holigarna caustica (Dennst.) for the management of pain. J. Phytomol. Pharmacol. 2022, 1, 37–46, doi:10.56717/JPP.2022.V01I01.005.
15. Xie, X.; Zou, J.; Wang, Q.-Y.; Shi, P.-Y. Targeting dengue virus NS4B protein for drug discovery. Antivir. Res. 2015, 118, 39–45, doi:10.1016/j.antiviral.2015.03.007.
16. Gasteiger E., Hoogland C., Gattiker A., Duvaud S., Wilkins M.R., Appel R.D., B.A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Humana Press, 2005, 571–607.
17. Geourjon, C.; Deléage, G. Sopma: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics 1995, 11, 681–684, doi:10.1093/bioinformatics/11.6.681.
18. Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: tools for motif discovery and searching. Nucleic acids research 2009, 37, W202-8, doi:10.1093/nar/gkp335.
19. Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J.E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858, doi:10.1038/nprot.2015.053.
20. Zhao, F.; Peng, J.; Debartolo, J.; Freed, K.F.; Sosnick, T.R.; Xu, J. A probabilistic and continuous model of protein conformational space for template-free modeling. J. Comput. Biol. 2010, 17, 783–798, doi:10.1089/cmb.2009.0235.
21. Sippl, M.J. Recognition of errors in three‐dimensional structures of proteins. Proteins: Structure, Function, and Bioinformatics 1993, 17, 355–362, doi:10.1002/prot.340170404.
22. Laskowski, R.A.; MacArthur, M.W.; Moss, D.S.; Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 1993, 26, 283–291, doi:10.1107/s0021889892009944.
23. Eisenberg, D.; Lüthy, R.; Bowie, J.U. VERIFY3D: Assessment of protein models with three-dimensional profiles. Meth. Enzymol. 1997, 277, 396–404, doi:10.1016/S0076-6879(97)77022-8.
24. Lovell, S.C.; Davis, I.W.; Arendall, W.B.; De Bakker, P.I.W.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure validation by Cα geometry: φ,ψ and Cβ deviation. Proteins: Structure, Function and Genetics. 2003, 50, 437–450, doi:10.1002/prot.10286.
25. Colovos, C.; Yeates, T.O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci. 1993, 2, 1511–1519, doi:10.1002/pro.5560020916.
26. Sivakumar, K.; Balaji, S.; Gangaradhakrishnan In silico characterization of antifreeze proteins using computational tools and servers. In Proceedings of the Journal of Chemical Sciences; Springer, 2007, 119, 571–579.
27. Guruprasad, K.; Reddy, B.V.B.; Pandit, M.W. Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng. Des. Sel. 1990, 4, 155–161, doi:10.1093/protein/4.2.155.
28. Pradeep, N. V; Vidyashree, K.G.; Lakshmi, P. In silico Characterization of Industrial Important Cellulases using Computational Tools. Adv. Life Sci. Technol. 2012, 4, 8–15.
29. Rogers, S.; Wells, R.; Rechsteiner, M. Amino acid sequences common to rapidly degraded proteins: The PEST hypothesis. Science 1986, 234, 364–368, doi:10.1126/science.2876518.
This work is licensed under the Creative Commons Attribution 4.0 License (CC BY-NC 4.0).